Major Eukaroytic Cell Organelles:
Campbell
Animal Cell*
&
Campbell Plant
Cell*
Organelle* : any
of the specialized structures within a
cell that performs a specific function
(e.g., mitochondria, ribosomes, E.R.,
Golgi, chlorolast, nucleus, etc...).
History of cell biology lies in the
era 1940-1970 when the functional
anatomy of a cell's
interior was worked microscopically
and biochemically giving an integrated
view of a cell.
NUCLEUS... a double membrane bound
organelle.
1st described & named by Robert
Brown 1836 - stamens
of orchid cells
1st
chemical analysis of nuclein, an acid, was
by
Frederich
Meischer
1869 - [DNA]
phosphorus molecule from pus of human cells (white blood
cells).
-
Largest organelle (a montage)*
- average dia = 6um [max10 um], volume up to 40 um3 is about 8%
of cell volume.
-
- found in all eukaryote cells
(except mature
erythrocytes & sieve tubes cells of phloem)
- evolutionary
origin... a membrane "surrounded" an early
prokaryotes nucleoid
(genophore).
Origin of nucleus is not well established;
possibly via
an
invagination-like process, i.e.,?
a Mesosome* is a folded invagination in the
cell membrane of bacteria that are
produced by the chemical fixation techniques
used to prepare
samples for electron microscopy.

-
-
-
-
-
-
Components* of the nucleus - fig 6.8
(11e-overview of cell)
a. nuclear
envelope* - nucleus is a double membrane
bound organelle - SEM image
b. nuclear
pore complexes* = pore
structure* & computer models
Human np's
human
pore complex has about
2000/nucleus & some 450+ nucleoproteins
(30 diff kinds)
inquiry based experimentation helped establish
the role of and
functional diameter of
pores = 10 nm
NUCLEAR TRANSPORT*
c. chromatin* - the genetic stuff 'inside of' the
nucleus is...
DNA (5x10-12gm) complexed with
histone proteins & acidic nuclear proteins
heterochromatin
(condensed & inactive
- dark in EM's)
euchromatin
(less dense & active
- greyish in EM's) structural image*
- d. nucleolus* -
a dense spherical structure... the site of
rDNA genes which make rRNA.
in the
Human genome there are 5 chromosomes with
nucleolar rDNA genes.
e. nucleoplasm* - interior complex-phase of the nucleus
that contains...
enzymes,
RNA's, solutes, chromatin, etc... akin to
cytoplasm.
- Role of Nucleus
- site of
genetic information, control of cell divisions
& heredity
-
Chromosome locations*
and
functional locations
within nucleus*
-

-
-
-
-
Nuclear Transport
Experiments to Determine functional
Pore Sizes & Transport Mechanisms
1960's -
Carl
Feldherr
injects gold
particles in
unicellular amoeba's
TEM's
showed particles congregating at nuclear pores
within a minute;
within 10 min, gold particles were in
nucleoplasm
see a micrograph*
1970's - fluorescent
tagged proteins
- showed proteins of less than 60,000 MW as passable
1990's - How do proteins get in/out?
(including ribosomal proteins & rRNA of ribosome)
Ron
Laskey
- studied a nuclear transport protein...
nucleoplasmin
(a thermostable
acidic protein helps assemble nucleosomes & in
ribosome biogenesis)
He radioactively tagged nucleoplasmin
& used autoradiography* to follow its movement
Laskey
experiment* panel a - shows nucleoplasmin (head & tail regions) enters nucleus
and suggests protein has an aa sequence that helps mobility
panel b - where is signal in head or tail? - they split & tagged
tail entered nucleus, thus it holds
aa
sequence
panel c - where in the tail? cut tail into pieces
& spliced to a
non-nuclear cytoplasmic protein.
►►► result:
nucleoplasmin holds a 17 amino
acid sequence
that targets transport into nucleus
it is known as the NUCLEAR LOCALIZATION SEQUENCE
(SIGNAL) (NLS).
suggests a likely
mechanism* for
nuclear protein transport.
 skip
skip
Mitochondria...
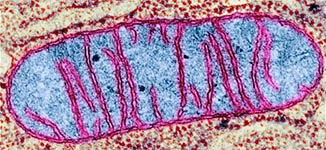
site of : Cellular Respiration - aerobic [redox
rx's] oxidation of
C6H12O6
--> CO2
+ H2O
-
Gas exchange in
cell - CO2 is released &
O2 is taken up & reduced to H2O
-
KREBS
cycle - an aerobic
pathway that oxidizes pyruvate
---> CO2 + H2O
- ETC chain, ATP
synthase & Oxidative Phosphorylation which
makes ATP
its role
: conversion of covalent bond energy in food
molecules --> into bond energy
in ATP
for cellular activities of all kinds by
coupling
redox transfers of e-
& H+ protons...
via ATP-synthase
--> to make ATP
Mitochondria
animation*
-
-

1st described
1857 by Albert von Kolliker & labeled as
'mitochondria' by Carl Brenda in 1898.
today: often
visualized via dyes*,
B&W-TEM*, &
false
color scanning SEM's*
-
double membrane bound organelle*
outer membrane - contains transport protein porin (passage of molecules up to 5K)
-
inner
membrane - very selectively permeable (i.e., impermeant to most molecules)
peri-mitochondrial
space* -
(in between) area where H+
accumulate (low pH)
cristae* -
inner membranes that hold the respiratory
assemblies of ETC (tomography*)
mitoplasm -
"matrix" interior compartment =
DNA, ribosomes, KC enzymes, etc
structure
: elongate cylinders to prolate
spheroids*
3-5 um long by 0.5-1.0 um dia,
-
"shape-shifters", mobile
-
number
: 20 to 1,000
per cell ; the more active
a cell = the greater their #'s
can make-up as much as 20% of cell's volume
contents:
has its own circular DNA - 16,569
nucleotide pairs: about 37
genes
has its
own ribosomes (prokaryotic size) &
protein synthesizing ability
holds enzymes for cellular respiration
-

-
CHLOROPLAST...
develops in the light from proplastids,
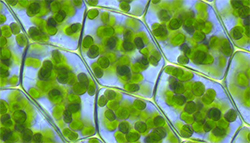
site*
of autotrophic metabolism = PHTS,
O2 evolution,
CO2
reduction to glucose
shape* - oblate spheroid in
higher plants, but shape is variable (stellate,
reticulate)
size: 2-3 um diameter by 5-10 um long
& numbering 15/20 - 100's/cell
-
Chloroplasm (stroma) contents = interior
compartment that holds
within itself...
1) internal membrane system made of thylakoid membranes:
Granal Stacks & Intergranal
Membranes*
2) 70s
ribosomes
(bacterial size) - Eukarya
have different size ribosomes [80s]
-
3) lipid droplets
4) 'naked' DNA pieces (circular, highly
super-coiled & repetitive)
-
5) pyrenoids (protein
centers for CO2 reduction) &
starch granules
6) enzymes of CO2 reduction to CH2O

What
is the origin of organelles as mitochondria and
chloroplasts?
are mitochondria &
chloroplasts endosymbionts*
by
Lynn Margulis - 1967
"Mitochondria
& Chloroplasts are derived
from archaean prokaryotes,
which were once free living, but joined into a
symbiotic relationship
with a primitive eukaryotic during cellular
evolution"

-

Some striking similarities of Prokaryotes with Mitochondria and Chloroplasts
that
support the endosymbiont hypothesis:
|
› both
organelles have double
membrane bound....
possibly the result of a phagocytotic engulfment?
of mitochondria?
of chloroplasts?
via eukaryotic evolution*
thus
a double membrane arose from endosymbiosis |
› both
are semiautonomous
:
derived from themselves, by divisional fission...
i.e., replicate independently
from their eukaryotic cell hosts |
› both have
their own DNA (a circular
molecule, like the DNA of prokaryotes)
& their protein
biosynthetic systems can make some of
own proteins
|
› DNA sequence
homology... each has
similar DNA sequences
mitochondria DNA related
to aerobic bacterial DNA
chloroplast DNA
related
to cyanobacterial DNA |
› ribosomes are same size
as bacterial
ribosomes* 70s vs. eukaryotes
with 80s sizes
[ S =
Svedberg
unitsG ]
Animation of
Endosymbiont Hypothesis(long
version)
|
 |
RIBOSOME...
... is a a non-membrane
bound organelle
... is a subcell ribonucleo-protein particle
(RNP) made of RNA & proteins
... discribed by George Palade in the 1940' via TEM
... is universal
to all known cells -
prokaryote & eukaryote
... is the cellular site of protein synthesis
(mRNA + ribosomes - artistic concept)
|
spheroid shape - 17 to 23 nm dia
(Noller model
& 2009 Nobel
Prize) |
| composed of 2 subunits |
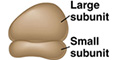 each contains rRNA & proteins each contains rRNA & proteins
|
small
subunit and a
large subunit, which binds tRNA's (Structure*)
prokaryotic & eukaryotic
composition = 35%
protein
and 65% rRNA
|
-
found in 3
different places* in cells...
-
1. free in cytoplasm, as individual
subunits or dimers,
-
2.
membrane
bound to the outer surface of Endoplasmic Reticulum membranes,
3. attached to
mRNA molecule in a
POLYSOME
[or
polyribosome*]
|

|
-
-
-
ENDOPLASMIC
RETICULUM... a set of membraneous
tubules contiguous with
nuclear membrane and found in ALL
EUKARYOTIC CELLS with a
nucleus,
(fig 6.11*)
making up 50% of
all internal membranes in eukaryotic cells.
The ER is involved in protein and lipid synthesis
in eukaryotic cells.
-
- Consists*
of flattened sheets,
sacs &
tubes of membranes making a
-
convoluted 3-D membrane network enclosing internal
spaces
- Lumen - is internal compartment of cisternae [makes up to 10% of cell's
volume]
-
-
2 Types:
Smooth E.R.
(SER
- tubular membranes without ribsosomes) &
Rough
E.R.*
(RER - surface of cisternae with
ribosomes)
Functions:
SER:
lipid & steroid
synthesis and drug
detoxification
(adds -OH's solubilizing them)
RER: synthesizes,
transports, & packages proteins into membrane vesicles
SIGNAL SEQUENCE*
- aa's @ N-term, bind, release into
lumen... Gunter Blobel
GLYCOSYLATION* -
adding carbohydrate groups to ER proteins
--->
glycoproteins
which will help
transport the proteins to specific cell sites

Golgi Apparatus*... is part of the
Endomembrane system...
a eukaryotic cell's
internal membrane system responsible
for
1. endocytosis - packaging of extracellular
molecules for internal distribution
2. exocytosis (secretion)
- packaging & delivery of newly synthesized
proteins/carbo's for extra-cellular secretion
Number &
image* of the Golgi
- up to 100 per cell
Size - 1 to 3 µm diameter
by 4 to 7 membranes stacks high
Structure
& function* - three parts
(or
sides)... |
 |
CIS side [entry side]... faces
R.E.R
proteins made on R.E.R.
pass from E.R.
lumen -->
vesicles --> cis Golgi |
MEDIAL cisternae elements...
proteins are modified with
sulfates, carbohydrates & lipids
modifications --> "address" vesicles to a
destination |
TRANS side [exit]... Golgi
side
modified vesicles leave as...
export vesicles, lysosomes,
other
membrane
bound vesicles |

|
|
Organelles of Cellular
Digestion...
LYSOSOME... a cytoplasmic single
membrane bound
vesicle,
derived via the ER and Golgi body
by glycosylation
changes, &
containing
hydrolytic enzymes with acid pH optima (pH 5.0);
Lysosome animation*...
function: intracellular
digestion via phagocyotosis* and autophagy.
Yoshinori Ohsumi
(2016 Nobel Prize for mechanisms of autophagy)
may have diverse
shapes, but mostly spherical*, with acidic
interior due to
lysosomal
membrane having ATP driven membrane H+pumpg
(faces inward*)
|
a sampling of
lysosomal enzymes involved in hydrolytic
digestion
|
| ENZYME |
SUBSTRATE |
|
acid phosphatase |
removes
phosphates |
|
acid nucleases |
digest
nucleic acids
|
|
proteases |
digest
proteins |
|
glucosidase |
digest
polysaccharides |
|
phospholipase |
phospholipids
& membranes
|
-
PROTEASOMES... large barrel shaped protein
complex, found in all eukaryotes and archaea,
and in some bacteria, that are responsible for intracellular Protein
Digestion... (structure)
ubiquitin
binds to proteins & transports them into a
proteasome figure*

-
-
Endomembrane System
visualization*... migration
path via vesicular
transport.
Was first
proposed by George Palade in 1940's and is part of a cell's
compartmentation
-
with the outer nuclear
envelope connecting to the rough ER & smooth ER.
-
Vesicles
made in ER flow, as transport
vesicles,
to the Golgi for modifications,
& from the Golgi to an intracellular
location or extracellular release.
c10e
fig 6.15*
Exocytosis:
Golgi modifies the
molecular composition and metabolic function
-
of the endomembranes vesicles as they flow from ER
through the Golgi,
-
& in turn, pinches off vesicles that give rise
to lysosomes & vacuoles,
-
or the plasma membrane
can fuse with vesicles born in the ER and Golgi.
-
the result is the release of proteins via a
Secretory
protein pathway &
-
other products to the outside of the cell in
exocytosis.
Endocytosis:
external material is captured into a membrane
vesicular
endosome,
which can fuse
with a lysosome for
intracellular digestion.
2013 Nobel prize
for vesicular transport
Protein
Sorting*
- proteins bound for different destinations have
diff carbohydrate tags
which can be recognized by unique glycoprotein
receptors in the cell.
so far we've seen 3 ways
to tag proteins for transport:
Nuclear Localizing Signal, Signal
Sequence, &
Carbohydrate Tags
proteins
have many sorting signals - table of
signals

- .
.
-
cytoskeleton...
-
network of protein
fibers running throughout the cytoplasm of
all cells (prokaryotes
& eukaryotes)
that provides a cell
its shape and a basis for cellular transport (ex:
cytoplasmic streaming).
stained cytoskeleton* & TEM's*
&
fibroblast*,
The Cytoskeletal
Proteins*
that make up the Cytoskeletal Network
include...
1. microfilaments (actin
proteins)... 7
to 8nm dia & of indefinite
lengths
actin is a universal (from protists to verts)
contractile eukaryotic protein
makes up 5% of total cell protein,
F-actin is a linear filament of polymerized monomeric globular
proteins of G-actin*
each microfilament is made up of two
helical, interlaced strands of subunits
.
... G-actin is a "conserved" polypeptide of 375aa with an ATP
recognition site
3 types of G-actins: - alpha
actins of muscle cells
[actin + myosin]
... beta
& gamma actins make up cytoskeleton and
are involved in cell motility
... actin filaments form crosslink patterns*
and can change lengths.
-
... example of function - make
up microvilli
of epithelia*,
cellular
membrane
protrusions that increase the surface area of
cells.
some other actin
filament roles*

-
-
-
-
-
-
-
-
-
-
2. intermediate
filaments...
(8-12nm dia - some ex:
keratin, vimentin & lamin)
protein fibers
[rope-like] with an intermediate diameter
spans cytoplasm providing framework for
mechanical strength.
made from a heterogeneous family of filamentous
proteins (pic*)
3. microtubules... made of tubulin proteins (also highly conserved
evolutionarily).
alpha & beta tubulin subunts assemble in um
long filaments with a
21-25 nm dia.
[ MT's* & pic-Hela
cell tubulin ]
that form depending upon bound GTP/GDP action.-
MT's are made of repeating globular units of 2
different proteins
alpha & beta tubulin,
which assemble
& disassemble*
[ +growth-*]
-
the major cytoskeletal
proteins are universal in all
eukaryotic cells.
-
a hypothetical roles
of cytoskeleton in cell structure*
-

Additional Cytoskeletal
Elements and/or Organelles...
Centrosome*
: Microtubule Organizing Center found in most
eukaryotic cells from which
MTs emerge: site of MT nucleation; organizes Cilia & Flagella
and the mitotic
spindle. Animal cell centrosomes contain centrioles
- plant cells DO NOT.
Cilia and Flagella and
Cell movements:
animination of cilia
& flagella structure*
Flagella are
microtubule (MT) extensions (100-200um
singly or in pairs) projecting from
cells that can push or pull a cell through an
aqueous media (ex:
bacterial flagella)
Cilia*
are MT extensions
(0.25 um in length with hundreds on individual
cells)
that move move stiffly
like oars, to propel a cell or move fluid over
stationary cell.
structure*: both have same structure
- 9 MT doublets surrounding 2 singlet MT's in
center,
collectively called an axoneme;
covered by plasma
membrane,
& often held by
cross-linking proteins (blue)
Basal Body* is a centriole found at the base of flagella or cilia
(anchor of cilia & flagella)

In 1965 Ian
Gibbons described a new protein studding
the length of each MT
doublet
in flagella, naming it
dynein (dyne for force and in
for protein)... which hydrolyzes
ATP.
The bending Motion is via Dynein arms* -
a
motor protein attaches & releases to MT
doublets,
and converts
chemical energy of ATP into mechanical energy of
conformational movement.
Dynein & Kinesin are motor
proteins* of intracellular
movements - walking
along MT's
ex: Kinesin*- is a
dimeric
motor protein powered by ATP
hydrolysis that changes conformation
& converts chemical energy into
kinetic energy, moving toward + (positive end) of MTs.
role of kinesin
motor proteins by Ron Vale and their discovery.
kinesin & dynein
motor differences (retrograde &
anterograde transports*
animations of
unseeable biology
Flagella
& Cillia movements are due to motor proteins as DYNEINsview@home
if
cross-links are
present
(blue in fig), the MT's are held in place, then dynein causes
MT
doublets to "curve (bend past each
other)* in restrained movement
if no cross-linking
proteins - one foot of dynein
arm binds as other releases
allowing MT to
"walk
along" MT
as doublets "slide" past each other
unrestrained.
one of the MT doublets walks or
slides toward the body of the cell on dynein
feet,
pushing in neighbor MT doublet toward the tip of
the cilium.

|
|
Roles of Cytoskeletal Protein
Elements in
Cell Structure and Motility -
Structural support:
actin
filaments bear the
TENSION forces (wires) of
the cytoskeleton
microtubules (in fig) are the COMPRESSION units (rods) [tensegrity]
offer inner structural support for
organelles (ex: toy & bridge*).
|
.
|
Cell motility:
contractile
force of muscles*:
myosin & actin
(microfilament)
are motor proteins
that via repeated cycles of binding and
release = movement (CONTRACTION)
amoeba's crawl*: is via a
psuedopodia due to the assembly/disassembly
[animation]
of individual actin
subunits on microfilaments shifting
between sol/gel
phases
cytoplasmic
streaming*: in plant cells occurs via
actin/myosin interactions and sol/gel
transformations which
results in a circular flow of cytoplasm
around the cell.
cell's respond to
pressure* by building
& branching actin filaments

|
-
Intercellular
junctions...
-
Cell surface regions specialized for intercellular
contact =
MULTICELLULARITY
especially
prominent in Epitheial Cells
of animals
-
3 Major Functions -
1. impermeabilize areas
2.
adhereing junctions
3.
communication
animation about
intracellular junctions*2.5min
-
-
Tight Junctions*
- they impermeabilize regions.
they prevent leakage of materials between
epithelial cells (normal vs.
celiac disease*)...
Desmosome - an adhering junction -
(anchors cells together)
spot desmosome
- spot welds made with
keratin
& cadherin proteins
Gap Junctions
-
intercellular channels for communication
[dia circa = 0.2nm]
allows ions, electric impulses, etc... to pass
between image*
Plants have no intercellular junctions as above
due to polysaccharide cell walls, but do have
Plasmodesma* - cytoplasmic connections between plant
cell walls [dia= 70nm].
a consequence: makes these plant cells act
like a single cell with channels allows
-
exchange of semi-large molecules to pass through
plasmodesma.

the plant
VACUOLE* animation*
is a Vacuolar
[tonoplast] membrane-bound sac
that plays roles in
intracellular digestion and the release of
cellular waste products
present in all plant, fungal, animal, &
bacterial cells.
In
animal cells, vacuoles are generally
small.
Plant cell lack
lysosomes.
Plant
vacuoles accumulate toxic wastes: phenolics, acids, and a
range of nitrogenous wastes
& water-soluble pigments, especially
anthocyanins -
responsible for red-pink-blue-purple
coloration in many (but not all) flowers and
fruits. Its interior is an
acid pH environment.
the (tonoplast)
vacuolar
membrane holds transport proteins, mostly
active-transport
carriers*
for one way accumulation of wastes and toxics into
the vacuolar spaces.
In plant
cells, vacuoles tend to be large
and play a role in maintaining turgor pressure*.
When a plant is well-watered, water collects in
cell vacuoles producing rigidity.
With insufficient water, pressure in the vacuole
is reduced and the plant wilts.
As plant
cells age.. onset of death is usually associated
with tonoplast leakage & breakdown.

 Key Concepts* Key Concepts*
-
all the links below
may enhance ones learning experience, but are not
required.
Tour of
a Cell5 min view at home
&
Virtual cell animationsrecommended
E. coli protein
molecular model &
comparison
prokaryotes & eukaryotes
Visual
Guide to human Cells & Inner
Life of a Cell & Scale of Biological parts
Molecular
animations to show how small molecules work
back
next lecture
copyright c2024
Last update - Feb 2024
Charles Mallery,
Biology 150, Department of Biology, U. of
Miami, Coral Gables, FL 33124
-
SKIP ALL THE MATERIAL BLOW THIS POINT
ignore the material below for Bil 150-pt
expanded table of differences between Prokaryotes
& Eukaryotes
Chaperones - proteins that
help fold other proteins into proper shape
(Sumanas protein folding
& degradation animation*)
Sumanas animation-vesicle
processing*
protein
recycling*
Targeting Signals for
semi-autonomous oragnelles (mito, chlp, peroxisome)
experiment
&
(model - eye lashes by P.Satir)
belt desmosome (zona
adherens) - wide band of desmosomes
MT's
might even play a role on consciousness ???
Skip the mateial below on this
page only
- Current Model of Nuclear
Pore Transport includes as many as 6 different
molecules including:
the molecules
an analogy to a moving
company
Importins
the delivery truck proteins
ATP &
ADP
the gas
GTP &
GDP
the unloading crew
and a
protein called Ran
the moving supervisor
an importin binds to
cytoplasmic protein with an NLS
(requires ATP)
figure
*
Ran + bound GDP complexes
with importin-cyto-protein & diffuses into nucleus
in nucleus GDP is phosphorylated & cytoplasmic
protein is released,
Ran escorts importin back to cytoplasm.
Exportins
- proteins found in nucleus that are counterpoints of
importins
RAN & GTP are also required, and a
Nuclear Export Signal
may be involved
= nucleosome &
chromatin packaging animation &
DNA supercoil
-
PLASTIDS...
-
group of double
membrane bound plant cell organelles...
found in all higher plants that produce all
the organics
required by metazoan cells [sucrose, etc...], and
store polymers of carbon and various pigments.
PROPLASTID...
a precursor plastid to all the other plant
plastids...
found in apical meristems* -
the dividing cells (≈
stem cells) of root/shoot tips
local cell environment defines
Type
of plastids*
to be made from proplastids...
etioplasts... chloroplasts developed in dark,
have an interior array of cystalline-
membranes & yellow-chlorophyll precursor-like
molecules
leucoplasts... non-pigmentous, 2x5 µm, variable
shaped plastids for storage
3 types: AMYLOPLASTS
(starch), ALEUROPLAST (protein),
ELAIOPLASTS (oils)
chromoplasts... plastids with water soluble
pigments, flower color, etc...
cilia:
move via alternating
power/recovery stroke cycles (ex: lining of windpipe & mucus)
moving fluids over a stationary cell; may beat up to 100 times per second.
Hypothesis of assembly of bacterial hair-like
pili bacterial movement via pili.
Role of Cilia: 1.
motion: as in clearing trachea of foreign
substances (fig)
2. stereocilia:
mechanosensing cilia of inner ear (fig*)
3. ependymal
cilia: keep Cerebral
Spinal Fluid flowing in the brain.
, i.e., "SIX-PACK MODEL"
made of a fibrillar protein network (claudins
& occludins) on apical side of epithelial.
gap junction - animation
Proteasomes &
Protein Digesting Drugs
- transport to + end
next
skip the material above
|



 each contains rRNA & proteins
each contains rRNA & proteins
