|
|
|
|
2 Fundamental Reaction
Mechanisms |
|
LIGHT
Reactions (photo-chemical
reactions... non-enzymatic)
molecular
excitation of chlorophyll*
porphyrin ring by light results
in...
charge
separation via hydrolysis
of
H2O
releasing
2H+ and ½O2
generation
of proton
motive force* (H+ gradient) across thylakoid
membranesx
synthesis
of ATP
via an ATP synthasex
reduction of NADP to
NADPH* within
Photosysystem I
|
DARK
Reactions (thermo-chemical
rxs [temperature
sensitive]...
thus
enzymatic)
CO2
fixation*
via (reduction)
reactions that are reverse of
glycolysis)
carboxylation rx: CO2 +
RuBP --> 2
PGA [ 1C + 5C
--> 2
(3C)
]
reduction PGA with NADPH
-->
PGAL (glycolytic-like)
regeneration of RuBP via
HMP (5C sugar)
pathway --> RuBP |
6CO2
+ 12H2O*
--> C6H12O6
+ 6H2O + 6O*2 |
 Both Light &
Dark reactions occur within the
Chloroplast - fig 14.29*
Both Light &
Dark reactions occur within the
Chloroplast - fig 14.29* |
Evolutionary origins of
Photosynthesis... |
The origin of
mitochondria & chloroplasts
may
have been symbiotic (ecb
1.20*)
|
Photosynthesis
is a redox process via the oxidation
of an electron donor
that
reduces NADP and creates a proton
motive force.
1st autotrophic
cells probably used H2S as donor e- donor: |
as the
purple-sulfur bacteria of
today use
CO2 +
2H2S --> CH2O
+ H2O +
2S |
cyanobacteria - are oxygenic
photosynthetic prokaryotes
CO2
+ 2H2O
-->
CH2O
+ H2O + O2
|
van
Neil equation
(1931)... Phts is
a light driven REDOX
reaction
CO2 + 2H2A
--> CH2O
+ H2O + 2A |
Evolutionary origin of
non-oxygenic photosynthesis is estimated
to be about 3.8 bya,
and geologists see signs of significant,
sustained levels of atmospheric O2
some 2.5 bya &
remains of cyanobacteria have turned up
in rocks 2.7 bya in Western Australia.
Traces of
RuBisCo,
the key phts enzyme that reduces CO2
have been found that are 2.7-2.9 by old. |
 |
|
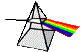
Capture of Light Energy:
Molecular
Excitation of Chlorophyll
Absorption of Light
Energy... physicists tell
us that light acts both as a
particle and a wave, depending
upon conditions:
Visible
light*: -
blue
light [440nm]
=
71.5 Kc/einstein
(1 mole
photons)
- near
red
[680
nm]
=
42.0 Kc/einstein (1
mole photons)
- red
light
[700nm]
=
40.9 Kc/einstein
(1 mole photons)
Chlorophyll
absorption spectra*
& Accessory
pigments absorption
spectra*
 phytol
tail*
embeds the porphyrin
ring (heme-like) in thylakoid
membranes
phytol
tail*
embeds the porphyrin
ring (heme-like) in thylakoid
membranes
has paired e's
with opposite spin which =
structural stability
Chlorophyll*
is in a Stable
Ground State
until light is
absorbed
light
absorption moves
non-bounded π
e's to
higher orbitals
1st excited singlet state...
2nd excited singlet
state...
[figure]
1st long-lived
state...

|
|
FATES of
Absorbed Light Energy:
1.
Re-radiated as vibrational
heat
2. Re-radiated as fluorescence* -
rapid emission of light of
longer wavelength & less
energetic
690nm
--> 740nm
in time frame 10-9sec,
3. Re-radiated
as
phosphorescence* -
delayed
emission of light much longer
wavelength
960nm
--> 980nm
much longer in real time
(1sec)
4. Induced
resonance transfer* -
vibrational e
excitation induces similar
electronic vibrations
in adjacent molecules causing
their excitation, etc...
5.
Photoionization* -
e's
enters into the photochemical
reactions of photosynthesis
excited electrons pass to an
acceptor leaving an ionized
chl+

|
|
Overview of the
Photosynthetic Light and
Dark Reactions*
The Light Reactions
are what
happens to the
photoionized
e-
of
chlorophyll.
PHOTOSYSTEMS*
- are
light
harvesting complexes of
chlorophylls,
Reaction
Centers,
accessory pigments, &
primary e-acceptors
in thylakoid membranes.
PS-II* - holds special
chlorophyll Reaction
Center PSII*
- water
cleaving part of PS-2 has 4 Mn atoms that oxidizes
& lose e-'s
► oxigenic photosynthesis was a great invention of
microbial
metabolism
PS-I* -
captures e- on
ferredoxin-(FeS) ►
Ferredoxin-NADP-reductase
► NADPH
► an
animation model of
the PS-I reaction
center from bacteria*
the
Pathway of e-
transport
between photosystems PS-II
& PS-I can
work in series
-->
Noncyclic
photophosphorylation
ecb
14.37*
in oxygenic photosynthesis
electron flow in along 3 legs:
from water to PSII, From PSII to
PSI, & from PSI to NADP+
PS-I
captures e-
into coenzyme NADP+
-> NADPH
PS-II releases
O2
from the splitting of H2O &
creates a proton gradient
or the Pathway of e-
flow may be only
through ONLY one photosystem
- PSI
Cyclic
photophosphorylation
(e flow)
ecb 14.37*
net result is synthesis of ATP
only with no NADPH: ►see how
mcb 12.42**
 light
rx summary animation (5 min)*view@home
light
rx summary animation (5 min)*view@home
|
Dark Reactions
of Photosynthesis...
occur in stroma (chloroplasm)
consume ATP
and NADPH
made in light reactions
reduce (fixation of) CO2
into CH2O
(sugars)
Mel Calvin used
radiocarbon tracing C-atoms during
dark reaction via 14CO2 "lollipop*"
& paper
chromatography* was
used to identify pathway.
1961 Nobel
3
different pathways are now known
to reduce C into carbohydrate in
plants...
1st. CALVIN
cycle or the
C3-pathway
one CO2
+ RuBP
(a 5C sugar) ---> two 3C sugars (PGA)
ecb 14.40*
two 3C sugars combine
---> 1 net
glucose
ecb
14.41*
Calvin
cycle is a reverse of reactions in
glycolysis*
Enzyme is
ribulose-bisphoshate
carboxylase/oxygenase
Rubisco EC
4.1.1.39 structure*
-
[50% of leaf
protein]
but Rubisco can oxygenate
(O2)
RuBP rather than carboxylate it...
--> Photo-respiration* -
[definition*] that
ultimately releases CO2.
This reaction
reduces efficiency of phts by 20-50% in
C3 plants.
To overcome plants make more
enzyme: 50% leaf protein is
'Rubisco'.
Genetically modified tobacco cells*
may improve
productivity by 40%.
 |
|
2nd. Hatch &
Slack Pathway... a 2nd Carbon
Reduction (fixation) pathway...
C4-pathway
uses
another
carboxylase enzyme
- PEP-carboxylase [EC 4.1.1.31]
14CO2
+
PEP*
--> OAA
[4C-acid] -->
malate in mesophyll cells
(H.Kortschak-1965)
a
4C-acid ---> 3C (PYR)
+ CO2
in mesophyll cells
and
CO2 is
reduced in Calvin Cycle of bundle sheath
cells to glucose.
C4-pathway*
& their differences in
leaf anatomy*
&
PEP-carboxylase (fig + fig)
Plants
that have only the Calvin cycle (C3
plants) have only RuBP
carboxylase, but
C4
plants have both Rubisco
& PEP carboxylase.
Several groups of crop plants (Corn,
Sugarcane, Sorghum), as well as
certain
dicots including
pigweed & halophytes, saltbush,
have developed C4 adaptations,
which allow CO2 uptake
& formation of a 4C molecule OAA
instead of the 3C PGA's
of the Calvin cycle.
KEY
comparison C3 vs. C4 enzyme
efficiency:
Rubisco's
- Km for CO2
is weaker in
C3
plants,
than PEP-carboxylase's Km
for CO2
is
in C4
plants (Km-graph*)
C4 phts
overcomes the tendency of
RuBisCO to use O2 rather than CO2 in
photorespiration by using a more efficient
enzyme to fix CO2 in mesophyll
cells
& shuttling this fixed carbon
into bundle-sheath cells, where RuBisCO is
isolated from atmospheric oxygen.

|
|
3rd. CAM
Pathway... (C4- Crassulacean
Acid Metabolism)
- CAM
plants...
are C4 plants
that don't separate C4
& C3
pathways in different parts of
leaf
(spatially)
but rather separate them in time
(temporally).
CAM was 1st studied in plant
family
Crassulaceae.
At night,
CAM
plants take in CO2
through open stomata at night,
when the succulents are in a
cool environment. The CO2
joins with PEP
to form the 4-carbon oxaloacetic
acid. OAA
is converted to 4-carbon malate
that accumulates at night in the
central
vacuole.
In the morning,
stomata close (thus conserving
moisture, as well as reducing the
inward diffusion of
oxygen). Accumulated malate
exits the vacuole and is broken
down to release CO2
that is taken up into the Calvin (C3)
cycle.
 Comparative
figure C4 v. CAM*
Comparative
figure C4 v. CAM*
|
These C4 temporal
and anatomical adaptations also
enable these plants to thrive in
conditions of (1) high
day temperatures, (2) intense
sunlight, & (3) low
soil moisture.
CAM plants have
same pathways & carboxylase
enzymes as in C4, within the
same cell...
thus show
TEMPORAL
not
SPATIAL
differences, regulated by
stomatal uptake.
CAM path
occur in a wide variety of plant
species, mainly in arid and tropical
regions...
other
examples of CAM plants: Crassulaceae
(Sedum,
Kalanchoe), Cactaceae (cacti),
Bromeliaceae
(pineapple) &
all
epiphytic
bromeliads including
Spanish moss,
Orchids,
and yucca. Also
included are weeds
as crabgrass
& Bermuda
grass
&
economically
important C4 crop plants include: corn,
sugarcane, & sorghum.
C3
plants represent only
about 85%
of the world's flora
C4
plants represent only
about
5%
of the world's flora (large # of
crops)
CAM plants represent
only about 10%
of the world's flora.
Comparative
figure C3 v.
C4 v. CAM*
 end of
material on Cellular Energy
Metabolism.
end of
material on Cellular Energy
Metabolism.
Up next is Molecular
Genetics with Dr. Daniel
DiResta.
|
LIGHT
close
close
corn, sugarcane, sorghum
SKIP THE MATERIAL BELOW:
OPTIONAL MATERIAL:
review
of Morphological basis...
Chloroplasts
&
their
structure - ecb
14.28
views
of detailed structure of
PSII and PSI
Karp
fig 6.10 PS-II
& Karp
fig 16.14 PSI
&
some
possible locales in
thylakoids
-->
combined
action of PSII & PSI
|