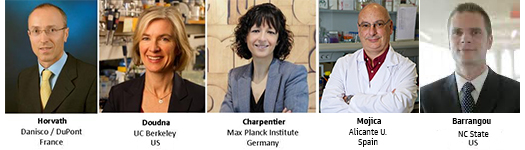
bacteria showed infections added new viral spacer DNA & their removal affected ability of yogurt bacteria
to resist infections. The viral DNA made RNA pieces that guided CAS nucleases to attack the invading
DNA and cut it up... 'a kind of bacterial immune system providing resistance to viral infections'.
CRISPR sequenes have subsequently been found throughout microbial life systems.
used to not only target viral DNA for cutting, but any DNA with a "guide sequence".
2011 - a. the essentials of how CRISPR works in bacteria are known and Emmanuelle Charpentier identifies an
essential RNA component 'tracrRNA of unknown function.
b. Eva Nogales, Doudna's colleague describes crystalline structure of the molecules
c. V. Siksnys (Vilnius U.) identifies a CAS-9 nuclease as the enzyme that cuts DNA
d. Jennifer Diudna & Emmanuelle Charpentier meet at conference in Puerto Rico and begin a collaboration
double strand cuts, a guide crRNA and a tracrRNA, which can be combined in a single transcript to
cleave any DNA target binding sequence with complementarity to a gene, offering a methodology based
on RNA-programmed Cas9 (that has considerable potential for gene-targeted editing applications).
culture. By end of 2013 Zeng's group had 64,571 unique CRISPR sequences for some 18,080 Human
genes (80% of the Human genome) with a 50% to 80% effectiveness.
method for genome editing.
are available to edit genes. Addgene ("the Amazon for Plasmids") is a non-profit that distributes
some 60,000 plasmids of various constructs (CRISPRs) to 20,000+ labs in 85 different countries for
researchers to use.
But, to date no CRISPR 'health' products to cure any illness are on the market. The US Patent Office
has 6,000+ CRISPR patent applications pending.
.