THey usied the JCVI-syn3A Mycoplasma cells with its minimum 493 genes. The computer simulation maps out the precise location and chemical characteristics of thousands of cellular components in 3D space at an atomic scale. It tracks how long it takes for these molecules to diffuse through the cell and encounter one another, what kinds of chemical reactions occur when they do, and how much energy is required for each step.
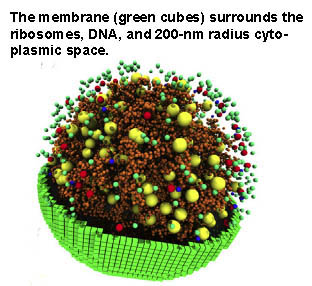

The model also was used to calculate the natural lifespan of messenger RNAs and to reveal a relationship between the rate at which lipids and membrane proteins were synthesized and changes in membrane surface area and cell volume.
The kinetic model opens a window on the inner workings of the cell, showing cell biologists how all of the components interact and change in response to internal and external cues.
Toolkit